Our Research
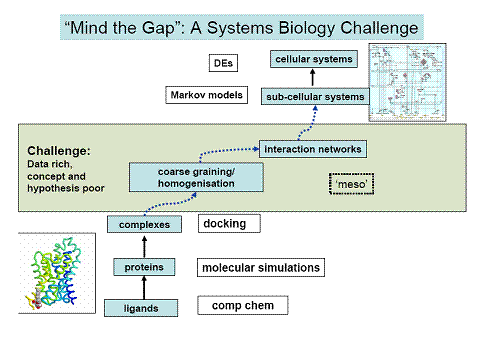
Biological function arises from the integration of processes interacting across a range of spatio-temporal scales. We aim to develop theoretical approaches which not only address problems at specific scales, but which bridge scales in a computationally efficient and mathematically tractable way. The approaches will be applied to a range of bacterial and eukaryotic sensory signalling networks, populated with data sets at different levels of completeness. The predictive models that we develop will be used to guide experiments, the results of which will be used to refine the models. This iteration between experiment and theory will both provide quantitative insights into important biological sysyems and enable the development of new tools with broad application.
Major advances in fast throughput technologies are producing cellular data in a quantity and format that both permits and demands their analysis by novel computational and physical methods.
Effective uptake of this approach in biosciences requires that modelling approaches should be anchored in biological experiment and generates the need for new mechanisms for mathematically and computationally handling data to produce meaningful insights into large-scale biological problems.
For details of major projects and approaches see pages using the links to the left of the page.